Beat Vögeli, PhD
Associate Professor

Contact Information:
Fax: (303) 724-1627
E-mail: [email protected]
Graduate Program Memberships:
NMR spectroscopy for the elucidation of conformation and communication networks within and between proteins, nucleic acids and small compounds
Recent breakthroughs in nuclear magnetic resonance (NMR) spectroscopy, X-ray crystallography, cryo-electron microscopy and other biophysical techniques have reshaped our understanding of dynamics and function of proteins and nucleic acids. We are beginning to appreciate a more holistic and fascinating reality where many, if not the majority, of the proteins and nucleic acids serve as interconnected communication hubs. Domain architecture is defined by the 3D average structure, but biomolecules are inherently dynamic entities. Our dynamic picture is still largely incomplete because no adequate methods are available to detect concerted conformational transitions at atomic resolution. These transitions, however, form the structural basis for relaying information from one molecular site to another (so-called allostery).
Despite impressive strides in artificial intelligence (AI) for predicting static, average macromolecular structures from sequences, AI cannot predict the dynamic fluctuations around these structures. However, visualization of these dynamics is essential to understand function, and NMR spectroscopy is the principal technique to answer fundamental questions addressing protein/nucleic acid dynamics at an atomic level. Using NMR, we contribute to the understanding of communication networks by modeling realistic ensembles of structures, identifying allosteric mechanisms, and exploring folding and interactions with other proteins, nucleic acids or small ligands. A cornerstone of our program is our newly developed exact nuclear Overhauser enhancement (eNOE) methodology for a realistic representation of molecular spatial sampling.
Exact NOE methodology
Our exact NOE (eNOE) approach marks a significant improvement over traditional NMR structure determination, which generally employs NOE rates in a semi-quantitative manner due to challenges like spin diffusion and low signal-to-noise ratios. The eNOE method allows for highly accurate distance measurements, facilitating a comprehensive structural representation via multi-state structural ensembles. To study domain interactions, we have extended the eNOE technique (individual distance restraints ≲8 Å) with electron spin resonance (EPR) for long-range distance measurements (≲25 Å).
We have developed a program that guides the analysis of eNOEs and extraction of distances for structure calculation (eNORA, available either in Matlab format or integrated into CYANA and Bruker DynamicsCenter). We apply the method to various proteins and also extend it to RNA and DNA.
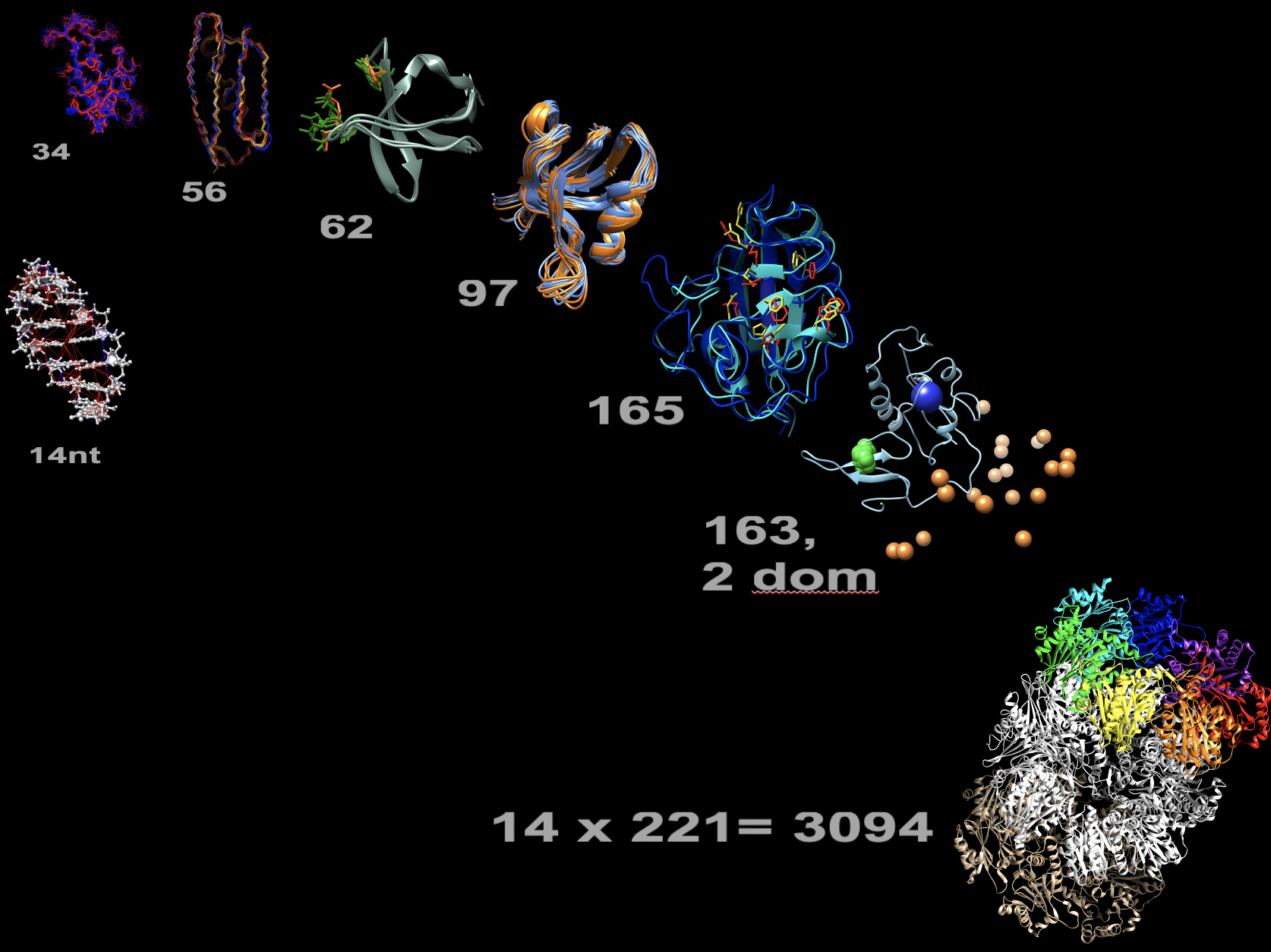
eNOE structure gallery. Proteins sorted by increasing residue length (grey numbers). Multi-state representations: WW domain of human Pin1, 3rd immunoglobulin binding domain of protein G (GB3), sarcoma homolog 3 (SH3) of chicken alpha-spectrin, PDZ2 of human tyrosine phosphatase 1E (hPTP1E), human cyclophilin A, two-domain full-length human Pin1, and 14-nucleotide UUCG RNA tetraloop (left). Only select distances: 20S core particle proteasome of T. acidophilium (bottom right).
Main Systems studied
Dynamics and allostery of Pin1
The human peptidyl-prolyl cis/trans isomerase Pin1 showcases complex allostery within its two-domain structure, significantly impacting biological processes such as mitosis regulation, Alzheimer’s disease prevention, hepatitis C infection, and cancer. Not only does the structure of the native WW domain dictate binding specificity, but its dynamic nature also facilitates information transmission to the distantly located catalytic site on the PPIase domain. Our findings indicate that Pin1's response to different ligands, which either strengthen or weaken domain interactions, triggers adaptive tuning of interdomain distances, thereby activating two separate allosteric pathways critical for allostery.
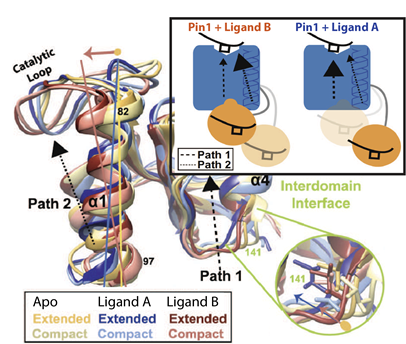
eNOE-based identification of distinct ligand-specific interdomain allosteric pathways in Pin1. Superimposed are two structural states of Pin1 for each the apo form, and complexes with ligands A and B. The large inset shows the resulting model: ligand B shifts the equilibrium of interchanging states towards a more compact form where allosteric pathways within the PPIase (blue) and WW domain (orange) are redistributed, whereas ligand A maintains conformations that favor the pathway inherent to apo form. Modified from Born et al, 2022, Nat Commun 13, 4546.
Functional disorder in intracellular transport
Despite its complexity, the human cell maintains precise intracellular distribution, a process crucial for understanding neurodevelopmental and degenerative diseases. Motor proteins such as kinesin and dynein navigate along microtubules to transport various cargoes, with many associated proteins displaying intrinsically disordered regions (IDRs). These IDRs often evade static structural representation by cryoEM or AI predictions, underscoring our project's goal to delve into underexplored structure-function relationships in cellular transport. Our research currently focuses on the following aspects of intracellular transport:
How can a single cytoplasmic dynein cover its entire functional spectrum?
There is only one major form of cytoplasmic dynein. Its specificities are partly due to a variety of cargo-specific motor adaptors, all of which contain IDRs, and the regulatory role of phosphorylation. We aim to reveal the mechanistic basis and functional implications of dynein specificity factors with a special focus on the dynein light intermediate chain (LIC), another IDR. We also elucidate the role of Pin1 (see above) in interactions between LIC and the dynein adaptors.
How does kinesin regulation by Olduvai protein impact human brain development?
Olduvai domains are encoded by Neuroblastoma Breaking Point Family proteins and show the largest human-specific increase in copy number of any protein coding region in the genome. This increase is linked to brain size growth, but also to schizophrenia and autism. To date, the molecular mechanisms underlying these functions are unknown. We aim to show how Olduvai domains, while being IDRs, impact kinesin transport. For these studies, we supplement NMR with other biophysical methods to probe structure, dynamics, and interactions of proteins involved in transport, and in vitro motility and cell-based assays to study resulting functions in collaboration with the J. Moore lab (Dept. of Cell & Developmental Biology).
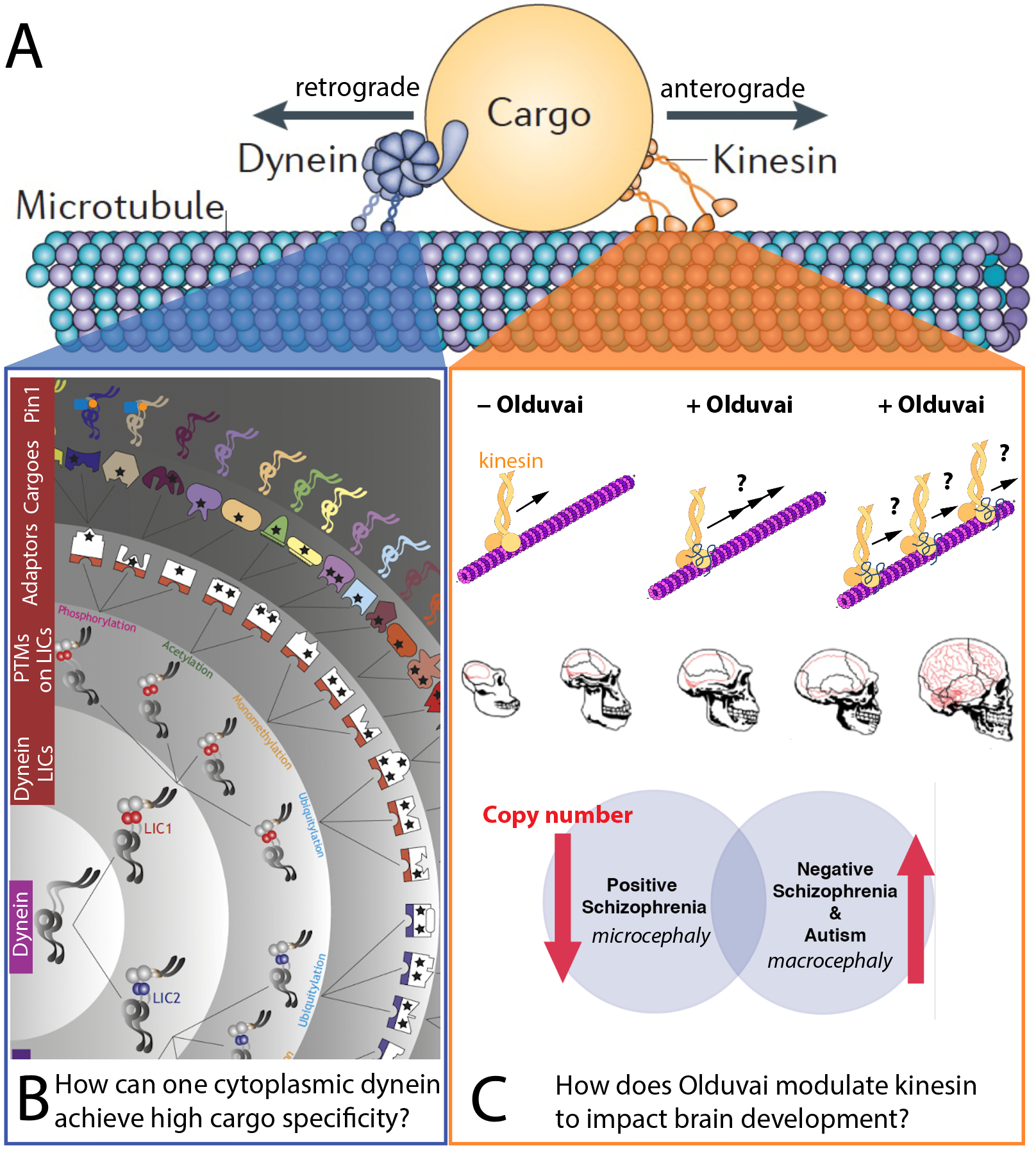
Knowledge gaps in microtubule-based intracellular transport (A). B) Our studies of dynein-driven transport address the astonishing functional spectrum of the single cytoplasmic dynein, presumably conveyed by various layers of specificity factors. C) Our investigation of kinesin-driven transport focuses on the impact of Olduvai protein on kinesin, which impacts human brain development. In part adapted from Kumari et al, 2021, J Cell Sci 134, jcs254870.
Immune regulation by Z-NA, ADAR1 and ZBP1
Z-RNA and Z-DNA introduce an unconventional left-handed helical structure, contrasting the typical right-handed formations of A- and B-form nucleic acids. Recent research has highlighted the importance of the historically elusive Z-RNA in modulating immune responses, with proteins containing Z-NA binding Zα domains, such as ZBP1 and ADAR1. ZBP1 acts as a sensor for regulating programmed cell death and inflammation. The double-stranded RNA editor ADAR1 is vital for preventing autoimmune responses by masking RNAs from innate immune sensors like MDA5 and ZBP1, thus averting erroneous immune activation. Our collaborative efforts with the Q. Vicens (University of Houston) focus on elucidating the structural and dynamic properties of Z-RNA and the associated proteins to understand their biological pathways and implications. See also the annual ABZ Meeting (abz.bio).
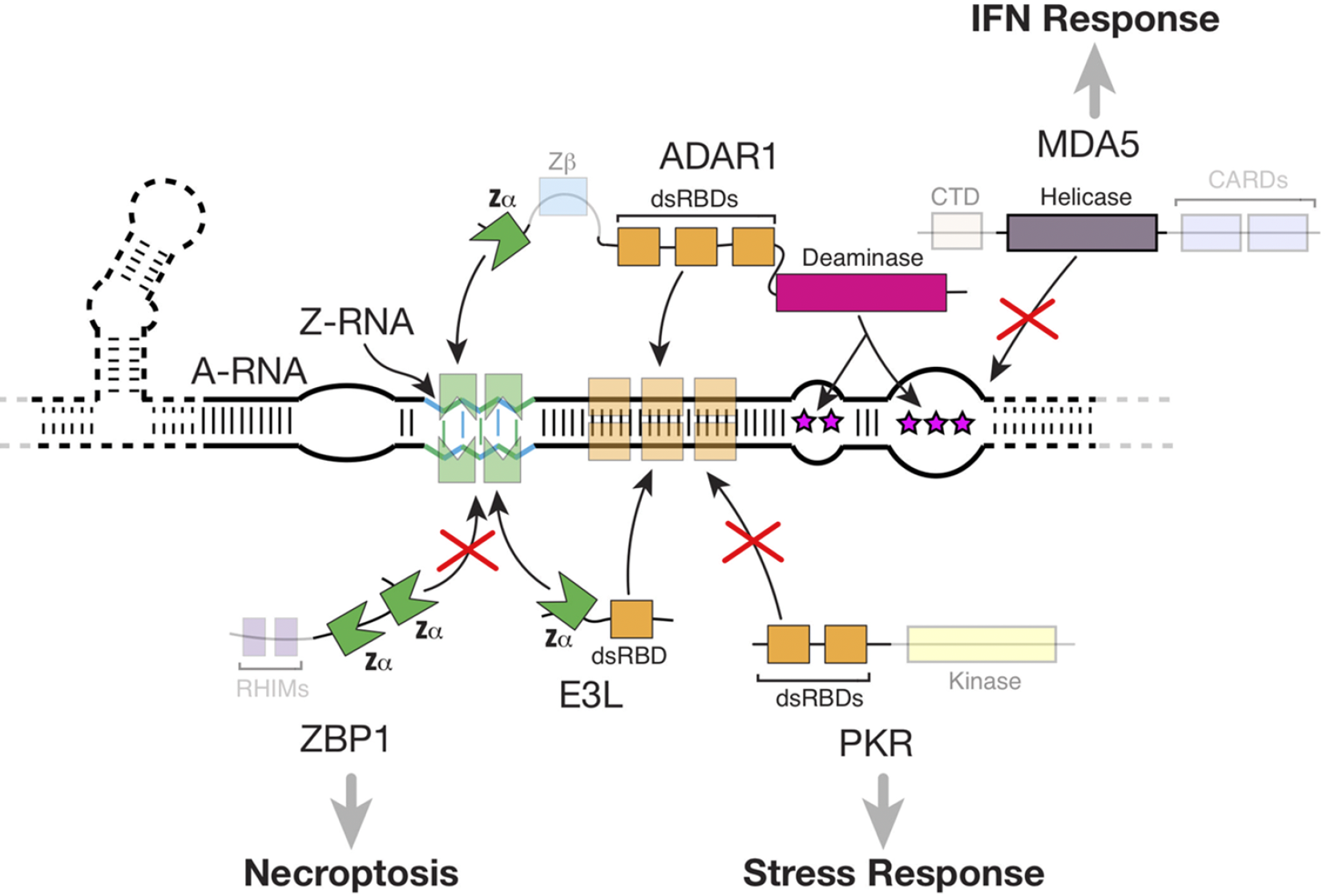
Interactions between Zα-containing proteins and Z-RNA. Pathways that depend on these interactions: direct shielding of dsRNA and Z-RNA by viral E3L as well as host ADAR1 proteins prevent recognition by ZBP1 and PKR, preventing activation of necroptosis and the stress response; editing of dsRNA by ADAR1 prevents MDA5 activation. Reproduced from Nichols et al, 2023, RNA 29, 273.
The role of structural dynamics in SASH1 function
SASH1, a tumor suppressor protein, plays a crucial role in various cancer types. Its diminished expression correlates with increased tumor growth, invasion, and metastasis. Interestingly, certain mutations in SASH1 can lead to premature hair graying and hyperpigmentation. Collaborating with the Y. Shellman lab (Dept of Dermatology), we explore the structure, dynamics, and interactions of SASH1, aiming to leverage this knowledge for targeted therapeutic interventions.
Ligand-protein interaction studies for drug design
Due to its high sensitivity, NMR is one of the leading techniques in the field of drug discovery. We partner with multiple laboratories to investigate ligand-protein interactions, advancing the development of therapeutic agents aimed at various diseases.

| Pictures | First Name | Last Name | Job Title | |
|---|---|---|---|---|
(1).jpg?sfvrsn=b79a16b4_0&MaxWidth=120&MaxHeight=160&ScaleUp=false&Quality=High&Method=ResizeFitToAreaArguments&Signature=7D988C3AB7992467EE9E02B1249723405FA0FA92) | Charles | Kroft | Graduate Student | [email protected] |
.jpg?sfvrsn=e4ac6b4_0&MaxWidth=120&MaxHeight=160&ScaleUp=false&Quality=High&Method=ResizeFitToAreaArguments&Signature=5911078FDE7ED9EDE3E0E9F50B96D193F5621FD2) | Morkos | Henen | Associate Research Professor | [email protected] |
 | Jeff | Krall | Graduate Student | [email protected] |
Pulse Sequence Code
Bruker pulse sequences for the measurement of dipolar cross-correlated relaxation between consecutive HN-N
Reference: B. Vögeli, Cross-correlated relaxation rates between protein backbone H-X dipolar interactions, 2017, J Biomol NMR, 67, 211-232
Bruker pulse sequences for the measurement of dipolar cross-correlated relaxation between consecutive HA-CA
Reference: B. Vögeli, Cross-correlated relaxation rates between protein backbone H-X dipolar interactions, 2017, J Biomol NMR, 67, 211-232
Bruker pulse sequences for the measurement of dipolar cross-correlated relaxation between intraresidual and sequential HN-N and HA-CA by MMQ
Reference: B. Vögeli, Cross-correlated relaxation rates between protein backbone H-X dipolar interactions, 2017, J Biomol NMR, 67, 211-232
Bruker pulse sequences for the measurement of dipolar cross-correlated relaxation between HN-N and HB-CB in Thr, Ile, Val
Reference: R.B. Fenwick & B. Vögeli, Detection of correlated protein backbone and side-chain angle fluctuations, 2017, ChemBioChem, 18, 2016-2021, DOI: 10.1002/cbic.201700312
Bruker pulse sequence for the measurement of eNOE buildups between methyl groups
Reference: C.N. Chi, D. Strotz, R. Riek & B. Vögeli, NOE-derived methyl distances from a 360 kDa proteasome complex, 2018, Chem Eur J, 24, 2270-2276, DOI: 10.1002/chem.201705551
Varian pulse sequence for the measurement of HA-HB(2,3) J-couplings
Reference: A. Born, M.A. Henen, P. Nichols, J. Wang, D.N. Jones, B. Vögeli, Efficient stereospecific Hβ2/3 NMR assignment strategy for mid-size proteins, 2018, Magnetochemistry, 4, 25, doi.org/10.3390/magnetochemistry4020025
Bruker pulse sequence for the measurement of eNOE buildups between all protons in a 15N/13C isotope-labeled protein via 3D [1H(F3),1H]-NOESY-[simultaneous 15N/13C(F2),1H(F1)]-HSQC
Reference: B. Vögeli, P. Güntert & R. Riek, Multiple-state ensemble structure determination from eNOE spectroscopy, 2013, Mol Phys, 111, 437-454, DOI: 10.1080/00268976.2012.728257
More Bruker pulse sequences can be found at:
http://n.ethz.ch/~bvoegeli/beat.html
Data Analysis Programs
eNORA2 MatLab code for the extraction of exact cross-relaxation rates and upper- and lower distance limits from NOESY buildups
Reference: D. Strotz, J. Orts, C. Chi, R. Riek, B. Vögeli, eNORA2 exact NOE analysis program, 2017, J Chem Theory Comput, DOI: 10.1021/acs.jctc.7b00436.
Example eNOE-Based Distance Extraction
From eNOE spectra to distances: a detailed protocol for converting assigned NOESY series peaks into exact distance restraints utilizing CYANA- or Matlab-eNORA
Reference: coming soon
The first comprehensive book on the fundamentals, analysis and applications of RDCs is out now!
Residual Dipolar Couplings: Principles and Applications

Royal Society of Chemistry book series New Developments in NMR, 402 pages
Residual dipolar couplings (RDCs) serve as prominent NMR tools for eliciting structural and dynamic details in both small compounds and complex macromolecules. Edited by Lishan Yao and Beat Vögeli, this book offers a comprehensive overview of RDCs, from fundamental principles to advanced applications in organic molecules and biomolecules. Renowned experts in the field delve into the latest advancements in RDC measurement and analysis.
Whether readers are seasoned professionals or aspiring students, this book equips them with a robust comprehension of RDC fundamentals and their potential integration into their research endeavors.