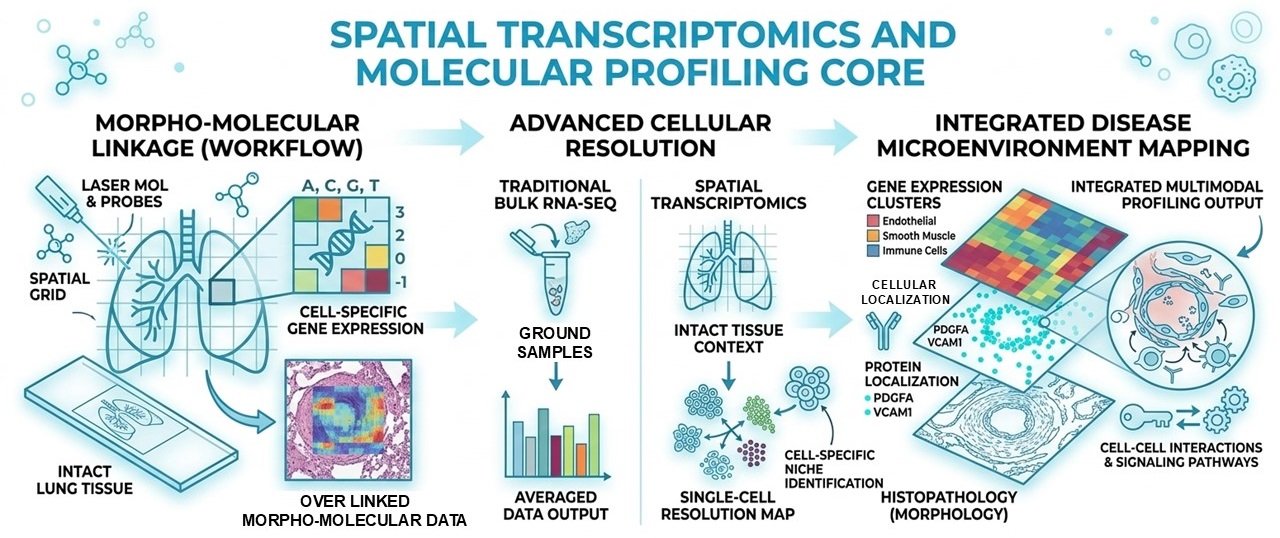
The Spatial Transcriptomics and Molecular Profiling Core within the Cardiovascular Pulmonary Research Laboratory (CVP) provides an integrated platform for studying the molecular and cellular architecture of pulmonary vascular disease. The core combines pulmonary pathology, spatial molecular profiling, advanced imaging, and bioinformatics to identify the molecular mechanisms that drive pulmonary vascular remodeling and inflammation.
By integrating modern spatial genomics with quantitative pathology and computational analysis, the core enables investigators to study gene expression, protein localization, and cellular interactions directly within intact lung tissue. These approaches allow researchers to map molecular signatures within specific vascular structures and disease lesions, providing insights that cannot be obtained through bulk or dissociated cell analyses.
This integrated framework supports projects investigating pulmonary hypertension and related cardiopulmonary diseases by linking molecular signals with histologic and structural features of pulmonary vascular remodeling.
Core Mission
The mission of the Spatial Transcriptomics Core is to provide investigators with the tools and expertise needed to:
- Characterize molecular signatures of pulmonary vascular remodeling
- Identify cellular niches within diseased pulmonary arteries
- Integrate histopathology with spatial transcriptomic and proteomic data
- Provide robust bioinformatic and statistical analysis pipelines
- Translate findings from experimental models to human pulmonary vascular disease
This multidisciplinary approach allows investigators to uncover molecular pathways driving pulmonary hypertension and inflammatory vascular remodeling, particularly within adventitial and immune cell niches of diseased pulmonary arteries.
Core Capabilities
The Spatial Transcriptomics Core provides a comprehensive set of services spanning tissue preparation, spatial molecular profiling, and computational analysis.
Lung Pathology and Tissue Processing
The core provides expert pathological analysis and tissue preparation for both human and experimental lung samples. Services include:
- Lung tissue processing and histologic preparation
- Quantitative assessment of pulmonary vascular remodeling
- Morphometric analysis of vascular lesions
- Multiplex immunohistochemistry and immunofluorescence
- High-resolution digital microscopy and image analysis
- Laser capture microdissection for cell-specific molecular analysis
These approaches allow investigators to correlate structural vascular changes with molecular and transcriptional signatures in pulmonary vascular disease.
Spatial Transcriptomics
The core provides spatial transcriptomic analysis of pulmonary vascular lesions using state-of-the-art technologies capable of resolving gene expression within intact tissue architecture.
Key capabilities include:
- Digital Spatial Profiling (DSP) for genome-wide expression analysis within defined microscopic regions of interest
- Single-cell spatial transcriptomics for mapping gene expression at cellular resolution within lung tissue
- Spatial analysis of pulmonary artery lesions, including plexiform, obliterative, and hypertrophic vascular lesions
- Construction of tissue microarrays from well-phenotyped human lung specimens
These technologies enable investigators to identify disease-specific molecular programs within pulmonary vascular niches, including fibroblast, macrophage, and endothelial cell populations involved in pulmonary hypertension.
Bioinformatics and Data Integration
The core provides an advanced bioinformatic and statistical pipeline for the analysis of spatial transcriptomic data.
Analytical approaches include:
- Dimensionality reduction and clustering of spatial transcriptomic datasets
- Differential gene expression analysis
- Pathway enrichment analysis (KEGG, GO, Reactome, Hallmark pathways)
- Integration of spatial transcriptomics with single-cell datasets
- Machine learning and trajectory analysis to identify disease progression pathways
- Correlation of molecular signatures with vascular pathology and morphometric measurements
These tools allow investigators to define cellular states, molecular pathways, and spatial niches involved in pulmonary vascular remodeling.
Technologies
The core integrates several advanced technologies to characterize pulmonary vascular disease.
Digital Spatial Profiling
Digital Spatial Profiling enables genome-wide measurement of gene expression within microscopic tissue regions. This allows investigators to analyze transcriptional programs within specific pulmonary vascular lesions while preserving spatial context.
Single-Cell Spatial Transcriptomics
Single-cell spatial technologies allow researchers to map gene expression at cellular resolution within histologic sections. These approaches enable detailed mapping of cell populations and molecular pathways within pulmonary vascular lesions.
Multiplex Imaging
Advanced multiplex immunohistochemistry and immunofluorescence enable simultaneous visualization of multiple proteins within the same tissue section, providing insight into cellular interactions and signaling pathways.
Laser Capture Microdissection
Laser capture microdissection allows precise isolation of specific vascular structures or cell populations for downstream genomic and proteomic analysis.
Together, these technologies enable multimodal analysis of pulmonary vascular disease at structural, cellular, and molecular levels.
Integration with Human and Experimental Models
The Spatial Transcriptomics Core supports studies across a range of experimental and clinical systems, including:
- Human pulmonary hypertension lung tissue samples
- Animal models of pulmonary hypertension
- Hypoxia-induced pulmonary vascular remodeling
- Schistosomiasis-associated pulmonary hypertension
- Pulmonary vascular disease associated with systemic disorders
By combining human tissue analysis with mechanistic animal models, the core enables investigators to determine how experimental findings translate to human disease.
Expertise and Leadership
The Spatial Transcriptomics Core brings together a multidisciplinary team with expertise in:
- Pulmonary vascular pathology
- Molecular and cellular biology
- Spatial genomics and transcriptomics
- Bioinformatics and computational biology
- Quantitative imaging and morphometry
The core is led by investigators with extensive experience in pulmonary vascular disease and spatial molecular profiling, including the molecular characterization of pulmonary arterial hypertension lesions and large-scale genomic analyses of lung tissue.
Supporting Collaborative Research
The Spatial Transcriptomics Core is designed to support investigators across the CVP research program and collaborating laboratories. By providing integrated pathology, spatial molecular profiling, and computational analysis, the core enables investigators to explore complex biological questions related to pulmonary vascular disease.
Through these capabilities, the core helps researchers:
- Identify novel molecular pathways driving pulmonary hypertension
- Map cellular interactions within vascular lesions
- Discover new therapeutic targets
- Integrate molecular findings with structural vascular pathology
This integrated approach is advancing the understanding of pulmonary vascular disease and enabling a new generation of spatially resolved molecular investigations.